atomium - a Python structure parser
Most scientific papers broadly take the form 'look at what we have just found out, and this is how we found it out'. That is, they report a piece of research, make a knowledge claim about the world, and explain their reasoning and the experiments they did to justify their claim.
This is not one of those papers. This paper belongs to the category of papers that report the creation of a tool that should make research easier. In this case the paper describes atomium - the Python library for parsing and processing macromolecular structures that I built.
Unless you're a structural biologist and/or programmer, that sentence might require some groundwork.
Python is a programming language - much of scientific research in Biology uses and relies on programming now for analysing data, simulating biochemical interactions, and generally automating tasks. Python has built-in modules for doing particular tasks, sometimes called libraries. For example there is a built-in library for searching text for patterns, and a built-in library for doing mathematical operations. Together these libraries are called the 'Python Standard Library' (yes both the individual modules and the collection are both called libraries) because when you install Python, you get these useful libraries bundled with it.
Then there are the so-called third-party libraries. These don't come as standard with Python, but are written by developers and made available for installation. These provide tools for fairly niche things that not every Python programmer might need. For example there is the requests library which provides easy to use tools for getting data from the internet, and the tqdm library for creating progress bars. These were written by people who don't work for the Python foundation and aren't affiliated with it, but who make these installable libraries available.
So, that's what I mean when I say I made a Python library. And 'parsing and processing macromolecular structures?'
Structural biology is the field of biology that deals with the three dimensional structure of proteins (a kind of macromolecule) - how their atoms are actually arranged. There are various experiments that scientists can do to work out the structure of a protein, and once they have, they have a set of coordinates for all the atoms in that protein. These are stored in a file, and can be opened by visualisation proteins such as PyMol to turn that list of coordinates into a rendering of it.
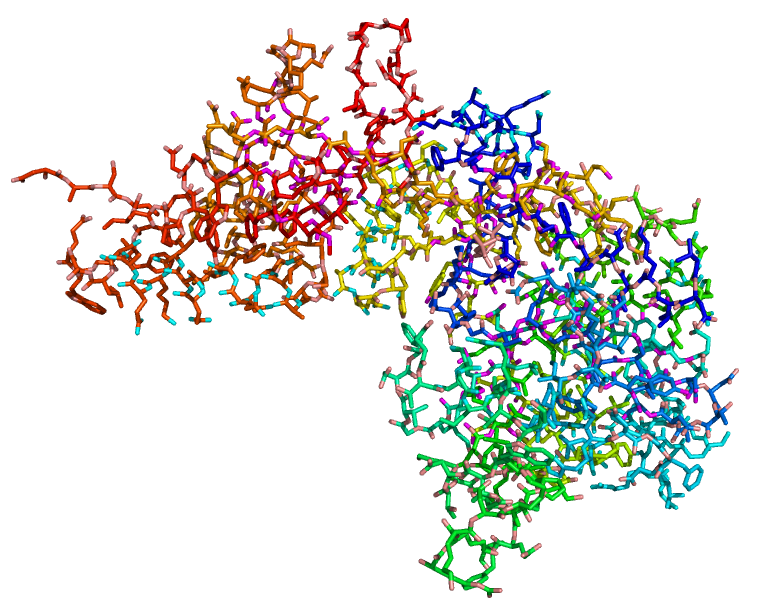
A visualisation of a protein's atom coordinates.
But if you want to actually analyse that structure - make calculations based on distances between atoms, compare algorithmically with other structures etc. - you need a tool to let you do that.
And that's what atomium is, and that's what this paper reports - atomium is a Python library for opening files containing the coordinates of proteins and other big molecules, parsing them into a data structure that represents the protein, and providing tools for doing common operations on them.
The full paper goes into detail about what it can do, how it works, and some key advantages over similar libraries that it has. For example, many structures need a certain amount of post-processing once parsed if the structure is to make any sense at all, and atomium is currently alone in doing this automatically.
Version 1.0.0 (the first stable release) was deployed in June 2019, and has received a few incremental updates since then. I think this tool is genuinely useful to the structural biology community, and I intend to maintain it indefinitely.