Projects
Here are the projects I have worked on in the past, or are currently working on - mostly software, mostly open source. You can filter them by the technologies they each use if needed, or click on them to get more details.
atomium
A lightweight, fast, modern parser of protein structure files.
Structural biology is the study of molecular structures - usually with hundreds or thousands of atoms. When analysed computationally they are stored in files such as the PDB file format, or mmCIF.
atomium parses these, and creates models of the molecules from them. These can be analysed, queried by atom type, rotated, transformed, or a variety of other operations.
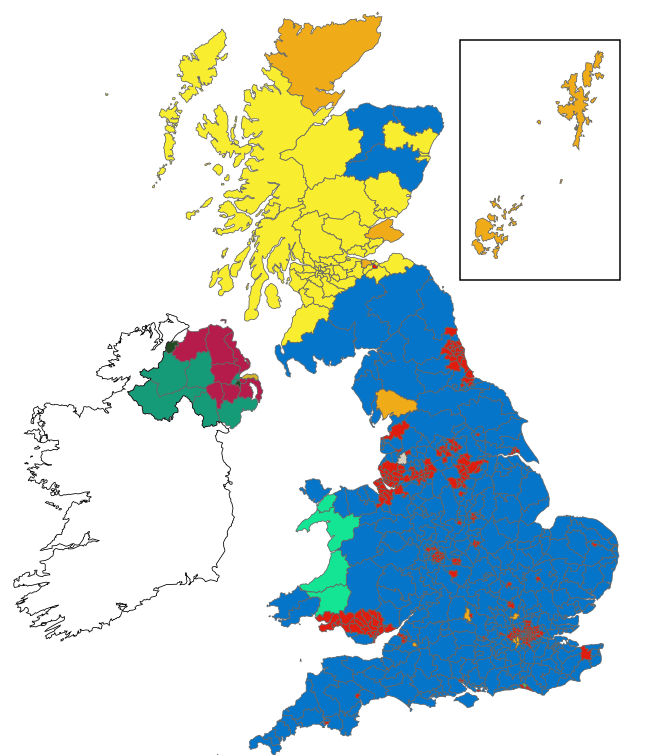
election19
A tool for running simulations of alternative versions of the 2019 UK general election.
election19 lets you define rules such as 'if 10% of Green voters had voted for Labour' and apply them to the result to see how this would affect individual constituencies, and therefore the overall election result.

nextflow.py
nextflow.py is a Python wrapper around the Nextflow pipeline framework. It lets you run Nextflow pipelines from Python code.
Nextflow is a pipeline workflow language for creating large, resource-intensive data processing pipelines, particularly in the life sciences. The pipelines are run from the command line.
This library is a Python wrapper around that, allowing the pipelines to be called from a Python script. It also provides abstractions for many of the outputs of these pipelines.
nextflow.py powers Goodwright's analysis platform, Flow.
molstar-react
A react library for adding the molstar protein viewer to a web app.
Molstar is a WebGL-based protein viewer, which can be used to display and interact with protein structures in a web browser. It is a very powerful tool, but it is also very complex, and so this library aims to make it easier to use.
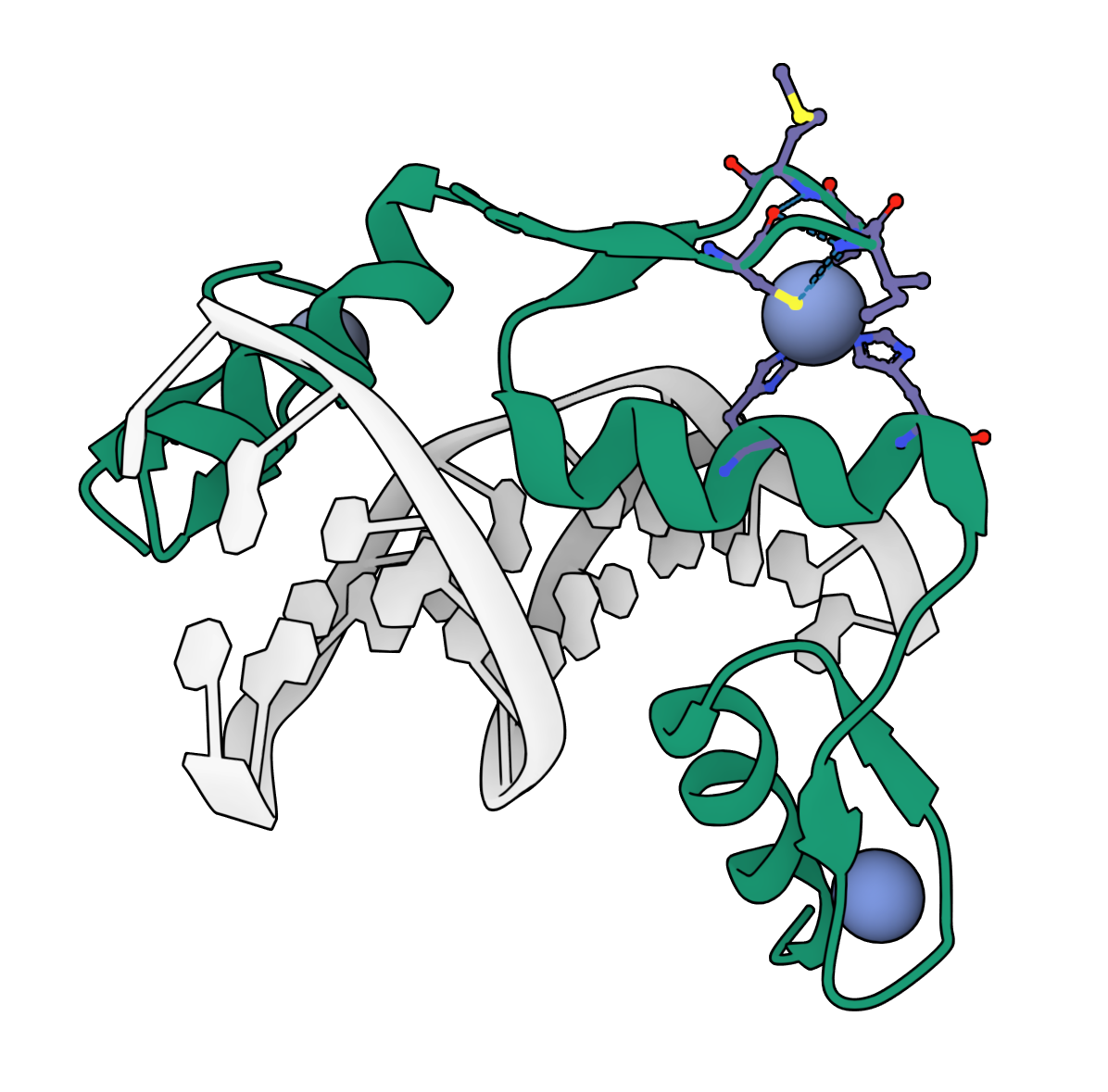
ZincBind
ZincBind is a database of zinc binding sites, automatically generated from the Protein Data Bank.
This project formed the core of my PhD research, which was an investigation into the properties of zinc binding sites in proteins, and how these might be used to train machine learning classifiers to predict them.
This resourcre contains all the identifiable zinc binding sites in known protein structures, as well as a prediction tool which uses the developed models to predict zinc binding in novel protein structures or sequences.

reddit-save
A Python tool for backing up your reddit saved/upvoted items locally.
Browsing through the things you've liked or saved on reddit can be a great way to recap stuff you once thought important. It is a personalised feed of posts and comments by the one person guaranteed to know what you like - past you.
However over time more and more of the older posts will be deleted or missing, and the historical record atrophies. Use this tool to back up those posts and comments to your computer where you can browse them offline, and where they are safe forever.
reddit-save will backup saved posts, saved comments, and upvoted posts. Crucially, when it is run again on the same location it will ignore any posts/comments previously archived - once something is saved, it's saved permanently.

Lytiko
A general purpose self-analytics app. It lets you track all kinds of things about yourself and tries to find correlations. It is currently being rebuilt as a comprehensive, end-to-end quantified-self platform.
Personal analytics is one of my core passions, and this app is the manifestation of that.

samireland.com
Originally a static website, then a PHP website, then a django templates site from 2016, and now built with wagtail (django) and next.js (react). Most of the things I have learned how to do in django were experimented with first on samireland.com.

kirjava
A Python client for accessing GraphQL APIs. Initially just a thin wrapper around the requests library, this is now a sophisticated but intuitive means of using Python to access modern GraphQL web services.
It also fully implements the GraphQL multipart request specification, allowing files to be uploaded with it.
pdbsearch
A library for searching for PDBs via the RCSB web services.
The RCSB is one of the organisations that maintains the central repository of all publicly available 3D protein structures. You can search through these via a complex advanced search page on their website, and the same functionality for searching structures is exposed via a web API.
This library is a Python wrapper around that API.

PetRank
A website created with Alex Harston to determine which pet is, objectively speaking, the best pet in the world.
The user is provided with two images of pets that other users have uploaded, and are invited to pick the 'best'. The chess ranking algorithm ELO is then used to turn these votes into a score, which is used to rank the animals.
The history of each pet is tracked and visualised.

django-random-id-model
This module provides a base class for Django models that gives them a random primary key id.
By default, database primary keys tend to just increment - the first will be 1, the second 2, etc.
This is not ideal - sometimes you want a random integer. This library handles that for you.

tiktok-save
A Python utility for backing up your liked and bookmarked videos on TikTok. It will download the videos themselves as mp4 files, and associated metadata for each video as JSON.
The rationale behind this, as with reddit-save, is to preserve content you have previously shown you like, without having to rely on the whims of whether a company continues to host it, or whether the uploader decided to delete it.
pygtop
A Python library that uses the Guide to Pharmacology API to present a Python interface to that data. The Guide is an expert-curated resource of pharmacological targets and the substances that act on them, and this library allows programmatic access to its data.
pdb2json
A simple web API for returning protein structures in JSON form. It uses atomium on the backend, and draws from any of the three atomium filetypes.
Protein structure files need complicated parsers to be able to access them, as they are stored in very specific file formats like .pdb or .cif. With pdb2json however, all you need is a browser and you can get those same structures in the universally understood JSON.

