ZincBind - The Database of Zinc Binding Sites
When we think of the living world, be it ourselves, plants, or anything else squishy and alive - we rarely think of metals. We think of ourselves as organic entities, and as metals as belonging to the cold, lifeless world of buildings and technology. Even if we did think of ourselves in purely chemical terms, we would usually think of the old classics like oxygen, carbon, and nitrogen.
But metals play a crucial role in every known living organism, including you and I. They might not account for much by weight, but there are metal atoms in all sorts of places within us. The oxygen in our blood relies on iron to be transported. Plants use magnesium to capture energy from the sun. And every thought you have ever had has relied on potassium to be conveyed along neurons. They are usually referred to as 'minerals' which is possibly why we don't think of them as metals, but they are everywhere and vital.
My PhD is all about one specific metal, zinc, and how it interacts with our proteins to give them abilities they would never otherwise have. Proteins are the chain-like molecules that do pretty much everything useful in our bodies. Produced from instructions in genes, they fold up into a particular 3D shape, and carry out their functions. I say 'chain-like' because they are chains of building blocks called amino acids, of which there are twenty, and these amino acids give proteins their unique abilities.
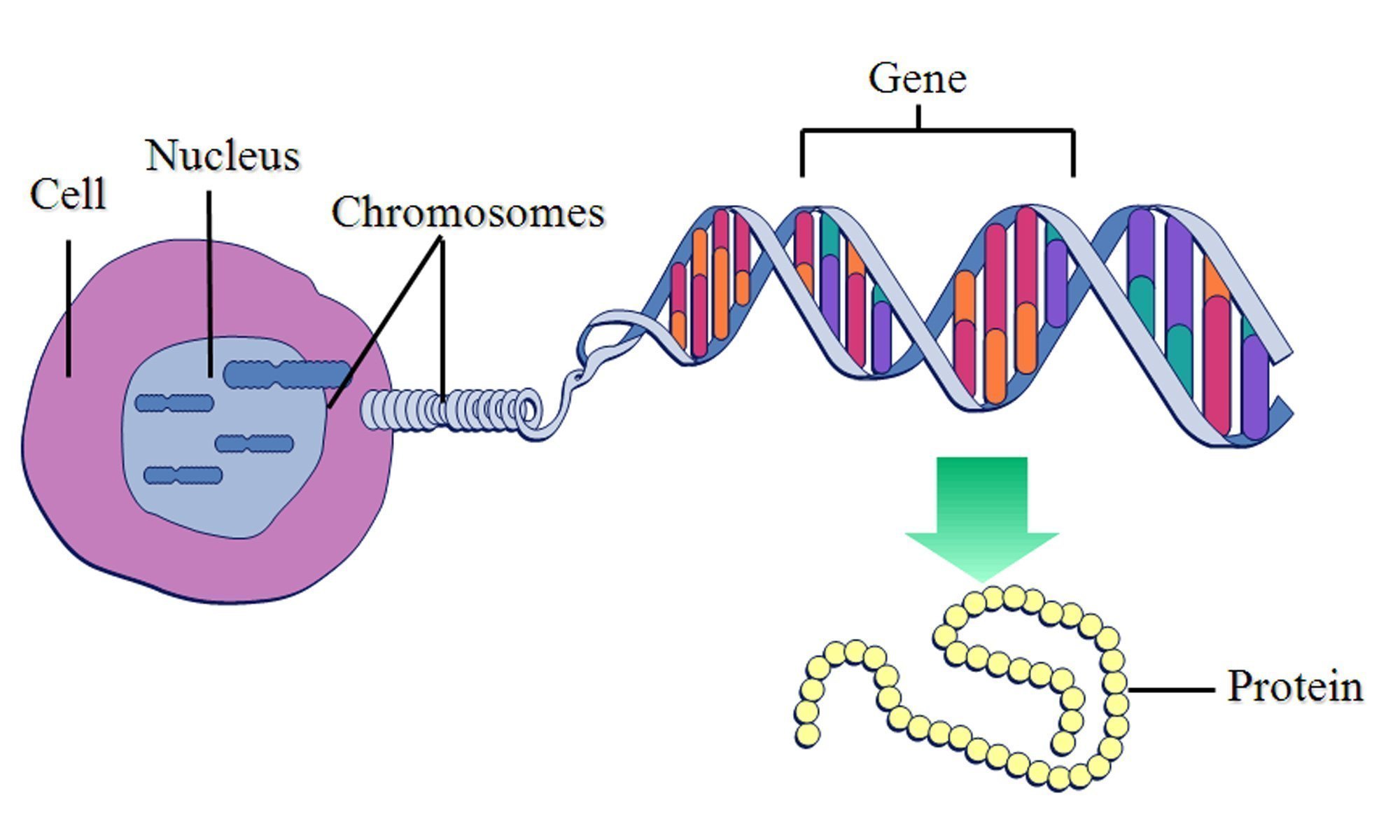
A diagram I stole from this study illustrating this principle.
For some tasks however, these twenty amino acids are not enough. They are all made of organic atoms, which are unable to do certain things which metal atoms can do. On a technical level, the electrons in a metal atom are arranged somewhat differently than in organic atoms, allowing them to interact with molecules slightly differently.
So, to access these abilities, proteins need to incorporate metal atoms. Since none of the amino acids have metal atoms in them, they have to do this in a rather roundabout way - they fold in such a way that three or four amino acids will cluster together to form a 'binding site' - a region of the protein which a metal atom will be attracted to. When the metal being drawn in is zinc, the area is a zinc binding site.
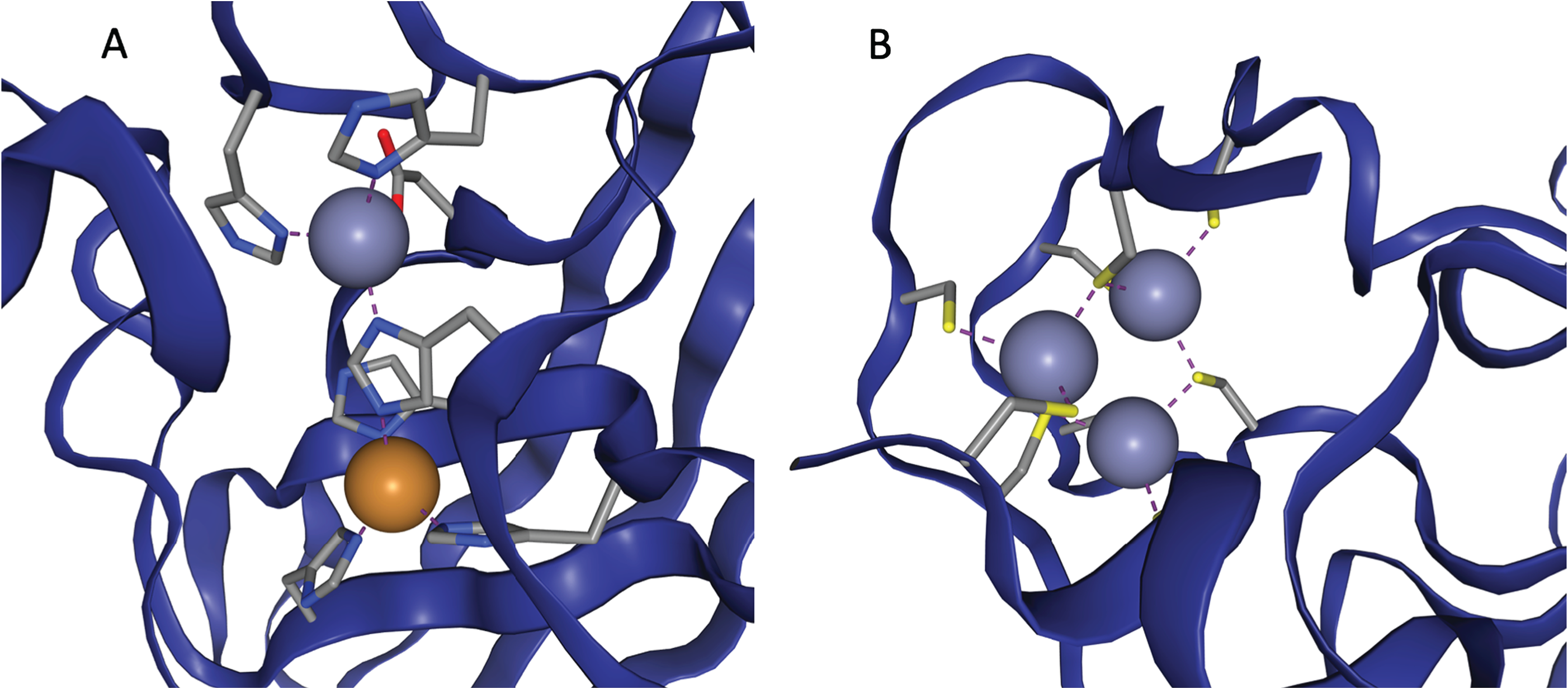
Two examples of zinc binding sites, taken from ZincBind.
Essentially, my PhD project is to find ways of predicting if a protein contains a zinc binding site, when you know either its protein sequence (i.e. the amino acids it contains and the order they are in, but not their actual arrangement in space) or its structure without zinc bound. After all, these zinc binding sites have certain characteristic properties, so it must be possible to use these to find them.
How do we know what these characteristic properties are? Well, we know the structures of many, many proteins by now - they are all stored in a resource called the Protein Data Bank. Many of these contain zinc, so there are thousands of zinc binding sites ready to be examined and studied.
That is why the first part of my PhD has been to create a single, definitive dataset of all known zinc binding sites, called ZincBind. I have created tools which go to the Protein Data Bank, work out which contain zinc atoms, looks at every structure, and gets all the zinc binding sites. This can be quite an involved process, as the software has to work out which amino acids are actually interacting with the zinc, work out whether any binding sites should be combined into a single site, and various other irritations that I won't go into.
The end results is a dataset of about 25,000 binding sites, from about 14,000 PDB structures. They are all annotated with the amino acids that the zincs stick to, the sequence of amino acids in the protein chain, information about the structure itself, and more. It will be regularly updated, and is the central dataset that my PhD project will use to create the predictive models I mentioned earlier.
That could well have been the end of it as far as my project is concerned - I only needed the dataset privately, and didn't need to do anything else to it. But access to the dataset has been made public via the ZincBind website. This is a Django web app which lets anyone access the data, query it, download it, and visualise each binding site in 3D thanks to the NGL plugin.
The ZincBind home page.
So that's ZincBind. The full paper goes into more technical detail about how the data was generated, and some of the preliminary analysis of the data that has already been performed, if you are interested. Otherwise, why not just explore the resource itself. After all, zinc is good for you.